An Introduction to Sequgio
Contents
2. Installing Sequgio
4. Other comments
4.2 Alternative counting method
1 Introduction
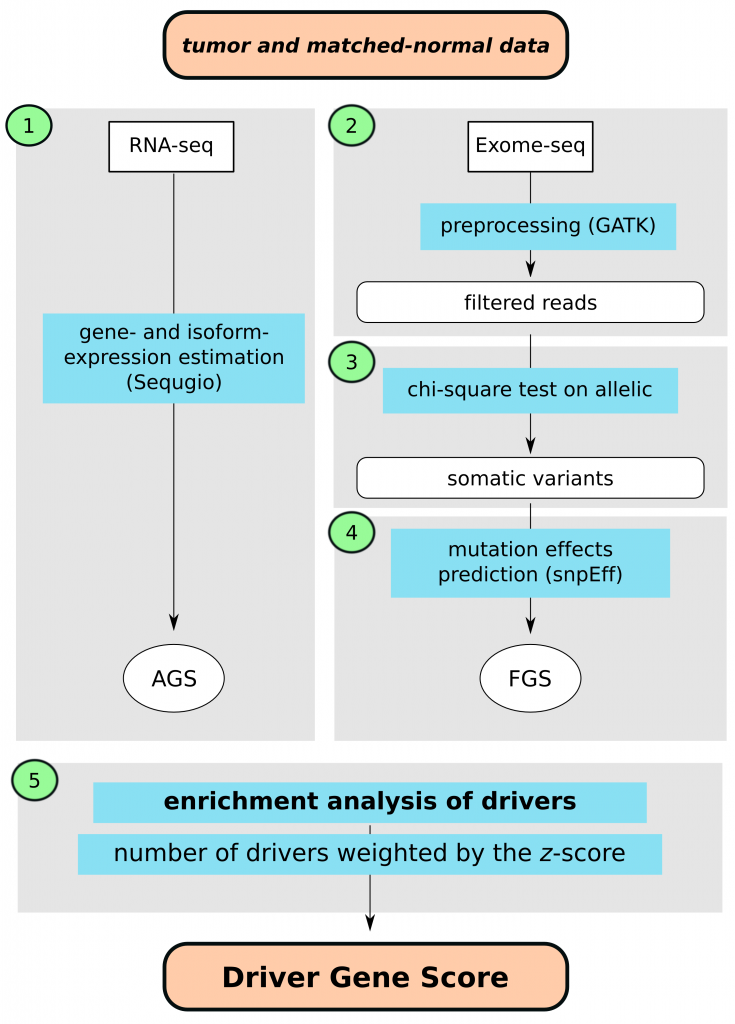
This document provides a brief guide to the Sequgio package, which is a package for gene isoform expression estimation, using a joint statistical model accounting for non-uniform isoform-specific read distribution (Suo et all. 2014). There are four components to this package:
- constructing annotation template txdb
- constructing design matrices used in expression estimation
- importing BAM files to create counts matrix
- estimating the expression levels, and optionally the read distribution and standard error of the expression estimates
Read intensity in the RNA-sequencing data is often not uniform, in which case the currently standard methods of gene and isoform expression estimates result in biased estimates. Sequgio accounts for non-uniform isoform-specific read distribution, yielding improved gene isoform expression estimation. A statistical regularization with L1 smoothing penalty is imposed to control the estimation. Also, for estimability reasons, the method uses information across samples from the same gene (Suo et all. 2014).
2 Installing Sequgio
Sequgio methods is implemented as an R package and will be soon available at bioconductor.org. In the meantime please use our developer version found on GitHub
Obtaining Sequgio package from GitHub (available now: 2015/01/20) From R execute below commands
install.packages("devtools")
library(devtools)
dev_mode(on=T)
install_github("Sequgio","Senbee",ref = "Stable")
dev_mode(on=F)
3 Example data
We show the functionality Sequgio package using RNA sequencing samples provided in the
RNAseqData.HNRNPC.bam.chr14. For demonstration purposes, Sequgio provides annotation template object (txdb, obtained in Step1) and design matrix (Design, obtained in Step2).
> library(RNAseqData.HNRNPC.bam.chr14)
> library("TxDb.Hsapiens.UCSC.hg19.knownGene")
> data ("TxDb")
> data ("Design")
For better performances the package supports parallel computing via the BiocParallel package which is loaded automatically. For parallel processing set the parameters to the ones suiting your platform. We will use sequencial computation here.
> param = SerialParam()
3.1 Step 1: creating annotation template (txdb)
The first step is providing TranscriptDb object that stores transcript annotation information. Here, we will use the one provided by the package TxDb.Hsapiens.UCSC.hg19.knownGene.
The TranscriptDb objects is then preprocessed to generate disjoint regions using the reshapeTxDb function. To obtain accurate results, set the read length parameter to match your experiment (72 nucleotides for the example data).
Further, since the example data contain only chromosome 14 we limit TranscriptDb to this chromosome as well (reducing computational burden).
# Example data
> seqs = seqnames(seqinfo(TxDb.Hsapiens.UCSC.hg19.knownGene))
> sel = rep(FALSE,length(seqs))
> names(sel) = seqs
> sel["chr14"] = TRUE
> isActiveSeq(TxDb.Hsapiens.UCSC.hg19.knownGene) = sel
> txdb = reshapeTxDb(TxDb.Hsapiens.UCSC.hg19.knownGene,probelen = 72L, mcpar=param)
# For the real data obtaining txdb could look like below
> txdb.ucsc = makeTranscriptDbFromGFF(file="pathway/to/genes.gft", format="gtf", dataSource=paste(http://cufflinks.cbcd.umd.edu/igenomes.html", sep="")), species="hg19.2014-06-02-13-47-56")
> seqs = seqnames(seqinfo(txdb.ucsc))
> sel = rep(TRUE, length(seqs))
> names(sel) = seas
> isActiveSeq(txdb.ucsc)=sel
> txdb = reshapeTxDb(txdb.ucss, probelen = 48L, mcpar=param_core)
This step has to be repeated only if the experimental parameters change, e.g. annotation database, reads length, with/without junctions etc.
If the user wants to focus on a subset of the “gene/features” in the genome, this can be done using the argument include.only. This takes a GRanges object as input. The purpose is to limit resources usage at BAM files input: this will be restricted only to the specific regions supplied.
# Let's consider only two genes for example
> GENES = genes(TxDb.Hsapiens.UCSC.hg19.knownGene)[c(1,3)]
> txdb = reshapeTxDb(TxDb.Hsapiens.UCSC.hg19.knownGene,probelen = 72L,
include.only=GENES, mcpar=param)
The ranges can be also be supplied later in the pipeline (see setCounts).
3.2 Step 2: creating design matrix
The second step is to create design matrices for each “transcriptional unit” (see references). These matrices will be used in the fitting procedure.
Several parameters can be tuned. The main one regards which kind of library is the experiment using: paired-end (“PE”) or single-end (“SE”, the default). This can be set with the method argument. The required mulen argument provides an estimate of the average fragment length.
Here we will use paired-end data.
> Design <- makeXmatrix(txdb, method="PE", mulen=155, sd=50, mcpar=param)
This step has to be only performed once, as step 1.
2.3 Step 3: importing BAM files and create counts matrix
Models are fit based on a matrix with read counts for every region in every sample. We will now import the aligned read counts in BAM files into Ras object called ’allCounts’. To do so we create a target object storing the filenames (with full path) and sample names to be used for count matrix headings. If BAI file are available, they can be provided in the target object.
The resulting object (allCounts) will count for every exons the overlapping reads.
Let’s first create the BamFileList object pointing to the BAM files. Data are pair-end so we set asMates to be TRUE.
> bflist <- BamFileList(RNAseqData.HNRNPC.bam.chr14_BAMFILES, asMates=TRUE)
Then we create a seqCounts object that will store some informations used in counting. A file name for file backed matrix can be provided or a random one will be generated. Also the root directory where the files are stored can be specified (defaults to working directory).
If the backing files (*.bin and *.desc) are then moved elsewhere, the files location informations in the seqCounts will have to be updated.
By default the function seqCounts will save the created object in the working directory, with the same fileName as *.bin and *.desc files. If this file is lost, it can be recreated allowing to link again to the *.bin file. To do specify seqCounts(…,existDesc=”fileName.desc”). A control on new exons/samples length will be performed but not on sample and exon names.
> seqObj <- setCounts(bflist,txdb,fileName="test")
If the user would like to reduce I/O resources usage when reading in BAM files, this can be restricted to specific region of interest specifying a set of ranges using includeOnly
#This would show if any subset is already set
> includeOnly(seqObj)
# Here we set it (see reshapeTxDb for GRanges definition example)
> includeOnly(seqObj) = GENES
# Or if we would like to remove it
> includeOnly(seqObj) = NULL
We then perform the actual counting with the function doCounts (no return value). This function saves the counts in the shared/filebacked big.matrix object.
> doCounts(seqObj,mcpar=param)
We can finally import in RAM the counts creating a standard R matrix.
> allCounts <- getCounts(seqObj)
In case the same big.matrix is used for more counting it should be reset to have counts zero.
To do so use the resetCounts function
> resetCounts(seqObj)
Again for ease of computation counts object is already provided.
> data(“Counts”)
We can see how many read we counted
> colSums(allCounts)
The counting procedure can be performed on single BAM files too or a subset of a BamFileList
For a single file we can proceed as follows
> bfl <- BamFile(RNAseqData.HNRNPC.bam.chr14_BAMFILES[1],asMates=TRUE)
> seqObj1 <- setCounts(bfl,txdb,fileName='test')
and then perform the following steps as described.
For a subset of a seqCounts we can proceed as described in the following code, using columns indicators (integers)
> doCounts(seqObj1,mcpar=param,which.sample=c(1L,4L))
or similarly using sample names
> doCounts(seqObj1,mcpar=param,which.sample=c('ERR127306','ERR127309'))
This last procedure would allow to have different nodes of an HPC platform to count a subset of the samples i parallel (each nodes get a different which.sample vector), while each node can use multiple cores to count individual samples. All the counts will be stored in the same shared matrix.
It also possible to subset a single sample (or many samples) based on genomic position. This can be achieved using filterBam function in Rsamtools package.
For example to read in the whole chr22 (in a human genome) we could proceed as follow
bfl <- BamFile(RNAseqData.HNRNPC.bam.chr14_BAMFILES[1],asMates=TRUE,
yieldSize=100000)
seqObj <- setCounts(bfl,txdb,fileName="test")
lenChr22 = seqlengths(txdb)['chr22']
bam.params = ScanBamParam(simpleCigar = FALSE, reverseComplement = FALSE,
what = c("qname", "qwidth", "mapq"),
flag = scanBamFlag(isPaired = TRUE, isUnmappedQuery = FALSE,
isDuplicate = FALSE,isNotPassingQualityControls = FALSE,
hasUnmappedMate = FALSE),
which = RangesList(chr22=IRanges(1,lnChr22))))
tmpFile = filterBam(bfl,tempfile(),param=bam.params)
doCounts(seqObj,mcpar=param)
2.3.1 Fixing QNAME
Some preprocessing pipelines deliver BAM files for paired-end reads where every mate has a slightly different QNAME. E.g. using the SRA toolkit to convert to fastq (fastq-dump), it will generate out two fastq files, and the QNAME in each of the fastq files will be appended with a “.1” for the first pairs and a “.2” for the second pairs. Similarly we might find that pairs have a different prefix.
The matching procedure implemented in Rsamtools requires mates to have the same QNAME, so a trimming is required. This can be achieved when declaring the BamFile or BamFileList object using the arguments ’qnamePrefixEnd’ or ’qnameSuffixStart’.
Unique qnames aren’t a problem in this sample file – just using it to demonstrate the usage.
First no trimming
> fl <- system.file("extdata", "ex1.bam", package="Rsamtools")
> param <- ScanBamParam(what="qname")
> bf <- BamFile(fl, asMates=TRUE)
> scanBam(bf, param=param)[[1]]$qname[1:3]
then trim prefix
> qnamePrefixEnd(bf) = "_"
> scanBam(bf, param=param)[[1]]$qname[1:3]
and now trim suffix also
> qnameSuffixStart(bf) = ":"
> scanBam(bf, param=param)[[1]]$qname[1:3]
In real situation the easiest approach is to specify trimming character directly in the call to BamFile or BamFileList.
For example in TCGA data (e.g. BRCA and LUNG) we have the following QNAME pattern for a pair of reads:
UNC14-SN744_125:6:2108:11845:164113/1
UNC14-SN744_125:6:2108:11845:164113/2
so we need to trim everything after (including) “/”
We would then define the BamFile object (or list if more than one) as follow
> bf <- BamFile(Filename,asMates=TRUE,qnameSuffixStart="/") ## single file
> bfl <- BamFileList(Filenames,asMates=TRUE,qnameSuffixStart="/") ## more than one BAM
## ALTERNATIVE WAY FOR STEP 3 WITH BASH-PYTHON (without fixing Qname) ##
This alternative method takes ~15 min=1 core=1 BAM file, and each BAM file can be counted in parallel (e.g. 12 samples = 12 parallel batch jobs)
1. Save your txdb into a text file.
> write.table(as.data.frame(txdb), "mytxdb.txt", sep="\t", row.names=FALSE, col.names=FALSE, quote=FALSE)
We already have pre-compiled TXDB for general purpose: /proj/b2012036/Dhany/Sequgio/txdb_hg19.txt and /proj/b2012036/Dhany/Sequgio/txdb_grch.txt
2. Write the list of annotated exon-pairs.
Do this, so that we can ignore novel exon pairs in this counting step. Make sure you still have the txdb variable in R:
> ex_list <- split(values(txdb@unlistData)$exon_name,values(txdb@unlistData)$tx_name)
> reg_vec <- sapply(split(values(txdb@unlistData)$region_id,values(txdb@unlistData)$tx_name),function(x) x[1])
> sizes_ex_list <- sapply(ex_list,length)
> n.exons <- sum((sizes_ex_list^2+sizes_ex_list)/2)
> exons.names <- unique(.Call("makeExNames",ex_list,reg_vec,as.integer(n.exons)))
> write.table(exons.names, "exons19.txt", row.names=FALSE, col.names=FALSE, quote=FALSE, sep="\t")
We already have pre-compiled list of annotated exon-pairs for general purpose: /proj/b2012036/Dhany/Sequgio/exons19.txt and /proj/b2012036/Dhany/Sequgio/exonsGrch.txt
3. Preview your merged BAM file.
Go to your bash command line, either open a new bash terminal or quit R. Find the pattern for the headers (i.e. first column) and answer the following questions:
(i) Can we tell which one is from pair 1 and 2? (0=no, 1=yes)
(ii) What’s the delimiter that separate headers and pair identifier?
(iii) Is the header on the left or right of the delimiter? (0=left, 1=right)
> samtools view -f 2 merged.bam | head
UNC17-SN1277:47:C0Y3YACXX:4:2205:18544:178539/1 81 chr1 10551 57 48M ...
UNC17-SN1277:47:C0Y3YACXX:4:2208:7653:169627/2 163 chr1 11193 54 48M ...
In the above example, (i) yes (=1) (ii) / (iii) left (=0). Thus, we define the “sep” parameter (separated by comma) as: sep=1,/,0
Beware that in some cases, the pair identifier is not /1 and /2. You must decide whether there’s separator or not.
4. List which chromosomes you want to count.
First, you need to know the chromosome names of the BAM files: 1,2,…,X,Y, or chr1,chr2,…,chrX,chrY. The names can be found on the second column of the BAM view (see above). Second, you need to mention which chromosomes you want to work with because there are other chromosomes in the BAM file, e.g. chrM, patch/putative chromosomes. To include those other chromosomes, you need to check whether your txdb recognizes those other chromosomes, too.
5. Run the Sequgio counting.
Run it in a BASH command line (still outside R). Specify the merged BAM file, chromosomes, txdb file, and output file. For >1 sample, it’s recommended to run in parallel (see Uppmax), not in serial. Never run many samples without batch/interactive session! Here is an example of counting for 1 sample:
> cd /proj/b2012036/TCGA/BRCA_BAM_files/
> bash /proj/b2012036/Dhany/Sequgio/newsequgiocount.sh file=UNCID_1120271.5969d3bb-19fb-4bda-9483-db4791257b95.sorted_genome_alignments.bam sep=1,/,0 chr=chr1-22,chrX,chrY txdb=/proj/b2012036/Dhany/Sequgio/txdb_hg19.txt output=UNCID_1120271.5969d3bb-19fb-4bda-9483-db4791257b95.sorted_genome_alignments.bam.txt
6. Join the counts of several samples
Several samples need to be counted so that you can continue to the next step. After joining the sample counts into 1 file, you can come back to R.
The syntax is: python makeAllcountMatrix.py [list of annotated exon-pairs (see no. 2)] [countfile_1] … [countfile_N] [samplename_1] … [samplename_N] > out.txt
> cd /proj/b2012036/TCGA/BRCA_BAM_files/
> python /proj/b2012036/Dhany/Sequgio/makeAllcountMatrix.py /proj/b2012036/Dhany/Sequgio/exons19.txt UNCID_1310380.5db1c37a-2329-4bd3-baf2-89d26a8959fa.sorted_genome_alignments.bam.txt UNCID_1311429.2efa4fd0-1c0b-49c8-a11f-606d8618ec97.sorted_genome_alignments.bam.txt UNCID_1313654.c14f7362-25c3-4e1a-acd7-871bb22ea8a3.sorted_genome_alignments.bam.txt UNCID_1314025.4543cecd-6333-473b-b5ce-96d33ab47706.sorted_genome_alignments.bam.txt UNCID_1338780.aaec0cbc-a493-48f4-9cc7-32629880ee24.sorted_genome_alignments.bam.txt UNCID_1338780.aaec0cbc-a493-48f4-9cc7-32629880ee24.sorted_genome_alignments.bam.txt UNCID_1367534.c7d2b685-5f11-4879-82d5-e983b5dbb07d.sorted_genome_alignments.bam.txt 1310380 1311429 1313654 1314025 1338780 1338780 1367534 > sample.counts
> allCounts
2.4 Step 4: fit models
Using the region(exons)-by-sample counts matrix (allCounts) and the design matrices object (Design) we can now fit models.
# model fitting
data(Counts)
data(Design)
iGenes <- names(Design)
# Fit a single transcriptional unit (one element in Design)
fit1 <- fitModels(iGenes[22],design=Design,counts=allCounts)
# More than one using list/for loops/mclapply/etc
fit2 <- lapply(iGenes[21:22],fitModels,design=Design,counts=allCounts)
# Fit all
fitAll<-bplapply(iGenes, function(x){fitModels(x,design=Design,counts=allCounts)},BPPARAM=param)
})
It is possible to get gene-level expression values adding up isoform estimates within gene. To do so proceed as follow
> sumByGene(fitAll,txdb)
where aFit is a list containing estimates (output of fitModels) and txdb is either a data.frame or a GRangesList (as returned by reshapeTxDb). The required columns are “gene_id” and “tx_name”.
References
1]Suo C, Calza S, Salim A and Pawitan Y (2014). Joint estimation of isoform expression and isoform-specific read distribution using multisample RNA-Seq data. Bioinformatics, 30, pages 506–513